Best tools for screening and reading research papers
 Blog
Blog Research
Research This post
This post
There’s a huge amount of relevant and high-quality research on any given subject, but researchers simply don’t have the time to absorb it all. It’s not just a question of discovering the best information, it’s understanding and applying it that matters. The risk is that important findings that could support new studies are being missed.To put this into context: there were more than 3 million articles published in 2018. And the problem isn’t limited to published research. The number of preprint servers - which are increasingly becoming an important resource for researchers - have grown by over 300% since 2015. Researchers typically spend 45 minutes reading a paper. And the average researcher reads around 250 papers a year. Trying to narrow down the literature that’s going to be most useful to you and then digesting all of that information are now tasks that are almost impossible without the help of technology.So, we’ve pulled together a short review of some of the most useful tools out there for searching and screening research papers. All of the tools presented here are either open-source (free to use and with open-licensed source code) and/or open-access (free to use).
Tools to build a repeatable search strategy
2Dsearch
At the heart of 2Dsearch is a graphical editor which lets you formulate search strategies using a visual framework where concepts are expressed as objects on a two-dimensional canvas. Search terms can be combined using Boolean operators into groups which can be expanded or collapsed to facilitate transparency and readability. This gives a more intuitive approach to search strategy development and validation. 2Dsearch consists of a query canvas and a search results pane, as shown below:
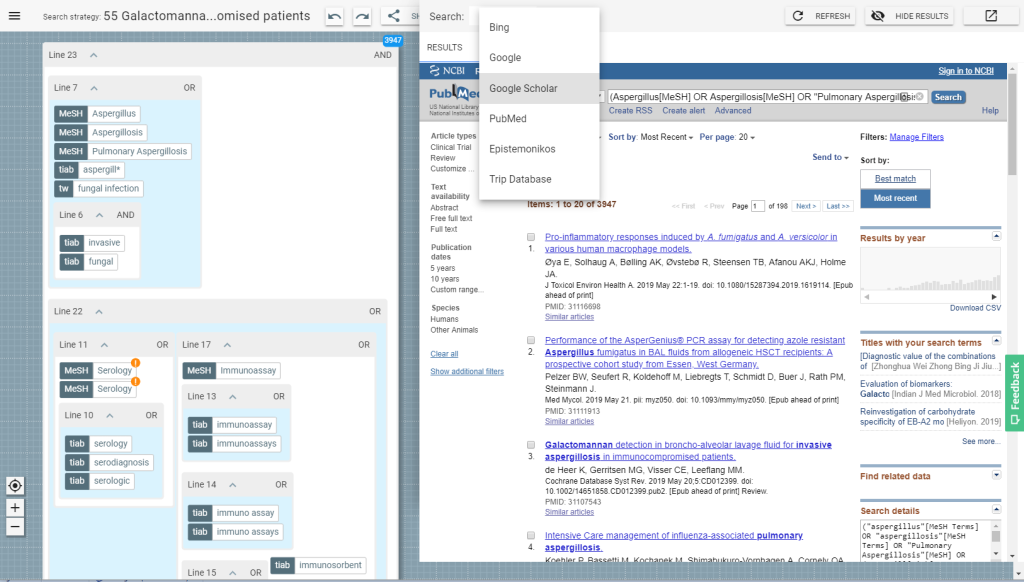
The application can take an existing search strategy and turn it into a visual structure. Take the following example:((telemedicine |telehealth |"Internet based"|telecare|"web based"|"mobile phone"|telemedical|videoconferencing|"text messaging"|"e mail"|telephone|"cell Phone"|pda|"e health")(diabetes|diabetic|insulin))| Telediabetes Although relatively simple, this query is still difficult to interpret, optimise or debug. However, when opened with 2Dsearch, its structure becomes much more transparent:
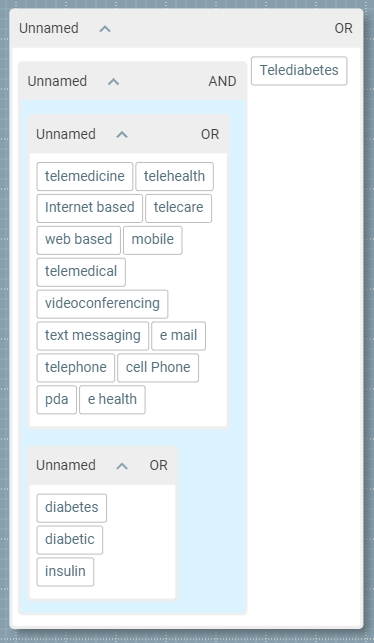
Visual search structure in 2DSearch
You can also use its automated search suggestions to identify and include related concepts:
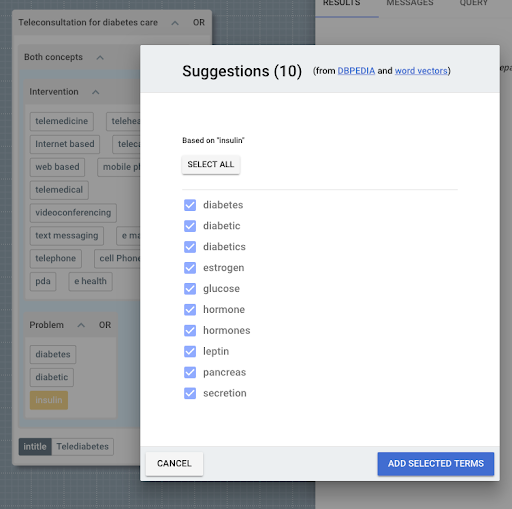
Automated search suggestions in 2DSearch
2Dsearch also provides support for automated translation to the syntax of other databases. For example, if you select the Query tab on the result pane, you’ll see our original query expressed as a Boolean string, along with a number of automated translations. Search strategies can be shared and exported as traditional Boolean strings.By offering new ways to for search strategies to be validated, shared and made reproducible, 2Dsearch can help academics search the world’s scientific literature in a systematic manner.
Tools for analysing search results
There’s a growing number of tools available for building and analysing citation networks. Here are just a few of the ones we’ve found to be effective when analysing a collection of articles.
Citation Gecko
Citation Gecko is an easy-to-use online tool that creates focused co-citation networks from seed papers. Seed papers can be added in the following ways:
- uploadinga BibTeX file
- connecting to your Zotero library
- connect to your Mendeley library
- select from search results
As you explore the cited-by and citing relationships, you can select these as new seed nodes to progressively build a citation map of the literature in your field

Similarity network tools make use of pre-built semantic indexes that are based on topic modelling or keyword co-occurrence. This allows semantically or thematically related papers to be clustered together. Search strategies can then be refined in a visual way which may reduce the cognitive load of filtering and reviewing a large number of search results.
Open Knowledge Maps (OKM)
OKM is an easy-to-use online tool that clusters search results according to semantic connections such as topics and keywords, which are automatically extracted from the source documents.Simply enter search term, choose a source repository (PubMed or BASE), and OKM will generate clusters that allow you to zoom in to explore semantically related papers.

Generated maps are given a persistent, shareable URL, and maps can be embedded on any web page.OKM helpfully shows which papers are Open Access via the unlocked padlock icon. When you select a paper in a cluster, OKM shows the article metadata and abstract in the margin, with a link to the full text article.

We can imagine a future tool that combines the functionality of Citation Gecko with OKM to show both citation and semantic relationships between papers.
Iris.ai
As with Citation Gecko, Iris.ai starts with a seed paper (via a public URL). It differs from other tools in that the input can also be a research question, although this requires a minimum of a 100-word description to be added for context.Similar to Open Knowledge Maps, Iris then clusters the results by keyword/concept, and as with OKM, it also generates a persistent URL that you can bookmark and share.

Search result clusters in Iris.ai
You can then drill down into each concept cluster to view semantically related papers. Selecting a paper allows you to then run a new search using that paper as the seed.
Connected Papers
Connected Papers is a new tool that also gives you a more visual overview of an academic field. Add a paper and it will create a graph for you of similar papers on the subject. It lets you build multiple graphs for relevant papers so that over time you can build up a visual understanding of the field you’re interested in, including trends, popular works and dynamics.
Tools for text mining and summarizing articles
Once you have filtered promising papers using the approaches described above, the next step might be to automate extraction of the knowledge they contain, to help give you a greater understanding of their findings and relevance.
GATE (General Architecture for Text Engineering)
The open-source GATE platform allows you to assemble a pipeline of prebuilt text mining components (or build your own) which you can run over a collection of documents. The output can be exported to various formats (XML, CSV) for later analysis.Some of the benefits of GATE include:
- no programming ability required to build your own text-mining pipeline
- comprehensive documentation
- online and in-person training courses
- the number and variety of plugins that enhance the out-of-the-box functionality
spaCy
Unlike GATE, spaCy does require programming skills in Python to use, although there are many tutorials online, including the excellent spaCy documentation at https://spacy.io/usage sciSpacy is a spaCy plugin that is useful for analysing biomedical papers, particularly for identifying concepts, abbreviations and negations.
Scholarcy
For those who don't want to download, install and learn new software, or don't want to write their own code, we built Scholarcy. Part of our goal was 'text mining for the rest of us': a simple way that anyone could upload a paper, or collection of papers, and return structured information such as keywords, key concepts, section summaries, facts, findings, and bibliographic references. Scholarcy provides a number of Application Programming Interfaces (APIs) to extract this type information at scale from a collection of research papers, speeding up analysis.Scholarcy also has a web application which renders the information extracted from a paper as a summary flashcard, that can be saved to a library for review, sharing, annotation and export.

Open Access Article - NPJ

Scholarcy Summary Flashcard
Unlike the original PDF, the summary card is responsive, and automatically resizes to fit your current device, so you can keep up to date with research wherever you are. The card shows the main concepts in the paper, linking them to their definitions in Wikipedia to provide the background knowledge needed to get the most from the article.The summary flashcard also highlights important facts and findings, as shown here in a summary of the introduction:

Expanded Summary Flashcard with important points highlighted
The study aims and findings are also summarised in the 'Highlights' tab, and the figures are extracted and cross-referenced in the text.

Scholarcy Highlights
For more advice and tools to help you read and write research papers, visit our blog.

